Becoming a Bioinformatician
13/11/2018
Let’s talk about the deep end. Specifically, being thrown into it.
Module 1 — Introduction to Bioinformatics using Perl
This first week has been a whirlwind. I have just started the Applied Bioinformatics MSc at Cranfield University. Within an hour of sitting down we’re facing the black screen and white text of the command line and being told to start programming.
Fortunately, we were given something between an introduction and a warning when we started; there are no essays and very few lectures, the aim of the game is to write code until you’re good at it, in multiple languages.
Our focus this week was Perl, the historical programming family used by biologists everywhere. After a brief outline of the main functions and data types we can use with it we began working through written practical booklets. These contained tasks designed to gradually build our base of knowledge on how to apply Perl in a relevant environment.
I was immediately engaged because it was clear that from the start that everything we learnt would be useful for our futures. These were not abstract challenges with no real-world application. Rather, by the time we received our assignment at the end of the week, we could build pipeline tools that can be utilised to uncover the sequence, structure and interaction networks of any protein required.
If there’s a corner to be cut (and more importantly, lines of code to be avoided) then I’m the first in line to find out how. Later in the week we were introduced to BioPerl. This is a collection of open source tools developed by the community that can be used as standalone modules or as part of a larger program.
Essentially, after writing scripts for a week, we were told that they’d already been written. Of course, without having first worked through our tasks we wouldn’t understand how the BioPerl packages were functioning and so it didn’t completely devalue our efforts.
Further, BioPerl is object-oriented, which felt like the next level. For Perl, we’re writing programs that process data they’re given and output a response. For BioPerl, however, we’re defining objects that interact with each other in a program. You can read this for a better explanation.
It was my first foray into coding; Perl is the first programming language I’ve tried to learn. I found that the learning curve is incredibly steep initially, but slowly you are brought around to a different way of thinking.
It’s frustratingly logical, and whilst Perl is supposed to be a more robust, flexible language (the unofficial motto is ‘there’s more than one way to do it’) compared to say, Python, the fact remains that a missing semi colon here, or open bracket there, will halt all progress. This is the case in any code you’re writing and so getting into the right habits has been a valuable lesson.
Although it appears that learning Perl may have more importance in understanding legacy code now, I have grown to enjoy using it. The Camel has always been the symbol of Perl and whilst the only official reason is its appearance on the cover of the first edition of Programming Perl by O’Reilly Media, it’s easy to see the parallels between the desert dweller and the often ugly, but always resilient programming language.
Categories & Tags:
Leave a comment on this post:
You might also like…
My Cranfield experience: How studying for the Strategic Marketing MSc landed me a job in my dream industry
For Shraddha Mahapatra, studying for a postgraduate master’s degree at Cranfield School of Management unlocked the path to a career working in her dream industry sector. Shraddha had gained an MBA in her native ...
Keen to develop your study skills?
Alongside the technical skills and academic knowledge that you will gain on your course, as a Cranfield student you have the opportunity to develop a range of other skills that can enhance your learning experience. ...
From classroom to reality: Supply chain insights from Cranfield’s Manchester study tour
Each year, Cranfield University organises a study tour for MSc Logistics and Procurement & Supply Chain Management students. For the 2025–2026 cohort, students were given the option to select one of three study groups: ...
Systematic literature review – Managing duplicates
One of the questions which often comes up when discussing the SLR process is how do I manage my references in the most efficient way during the process of going from my search results to ...
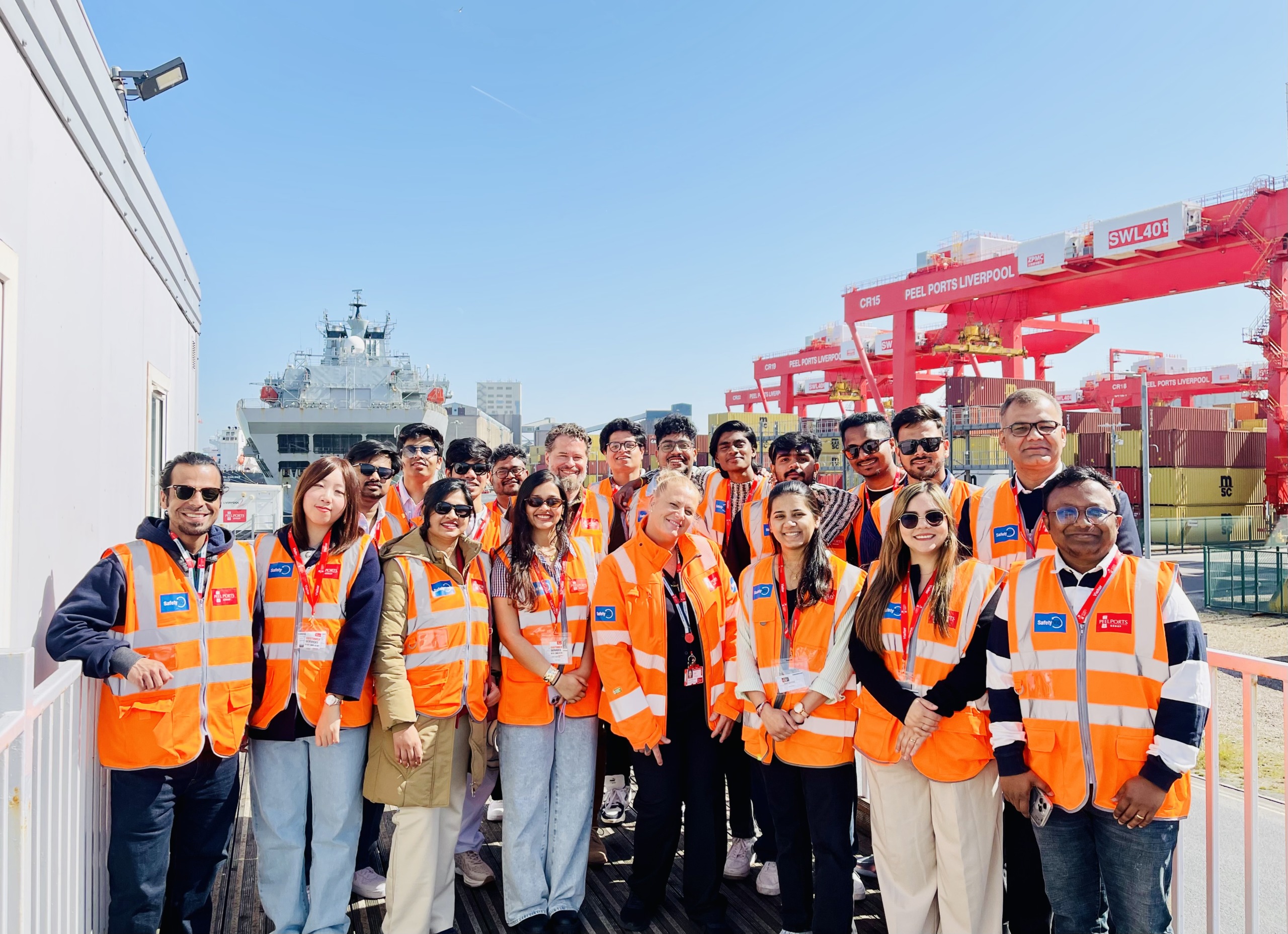
Liverpool study tour: Connecting classroom learning with industry practice
From 21 to 24 April 2026, the MSc Logistics and Supply Chain Management cohort at Cranfield University took part in a valuable Liverpool Study Tour. The visit was a strong example of our close ...
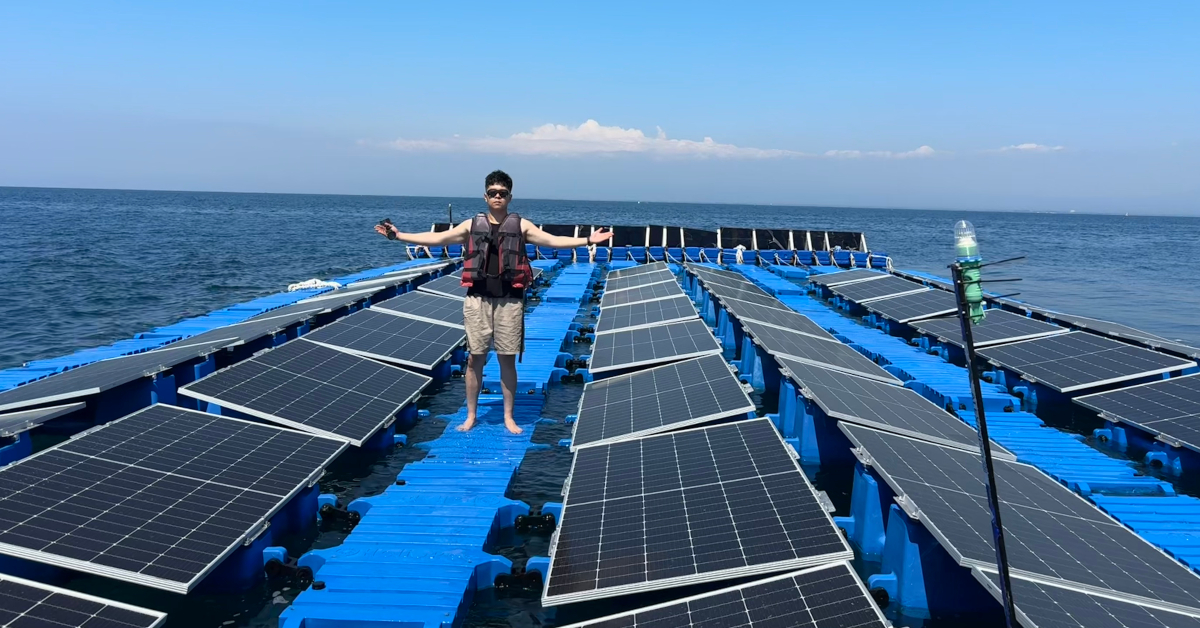
From wave tank to ocean: seeing my work come to life in Indonesia
Gili Ketapang is a small island in East Java, Indonesia. Around 2% of the population of Indonesia lives without access to electricity but the InnovateUK-funded Solar2Wave project aims to make sure 100% of the ...






